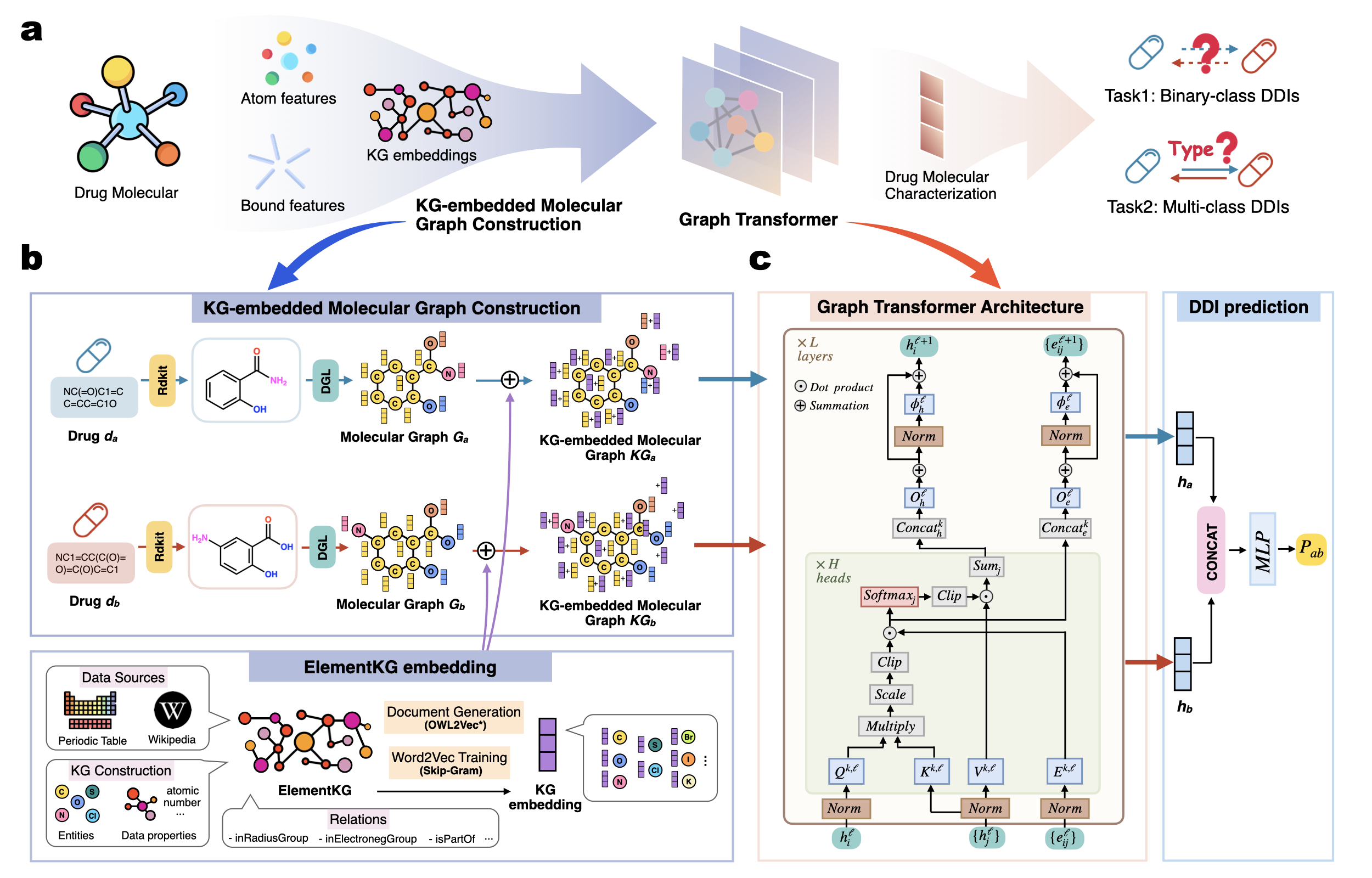
Identifying drug-drug interactions (DDIs) is a critical task in pharmaceutical research and clinical applications, as these interactions can pose serious medical risks. Deep learning models, known for their ability to accurately predict DDIs, have become powerful tools for enhancing prediction accuracy and efficiency. However, many existing approaches fail to fully incorporate chemical information and lack interpretability when exploring DDI mechanisms. In this work, we propose TRACE, a transformer-based graph representation learning framework that integrates chemical knowledge into DDI prediction. Extensive experiments demonstrate that TRACE outperforms state-of-the-art baseline models under both in-distribution and out-of-distribution settings, highlighting its strong predictive performance and generalization ability. In terms of interpretability, TRACE leverages its attention mechanism to effectively identify high-risk substructures that may trigger DDIs. In summary, TRACE not only provides new perspectives for elucidating the underlying causes of DDIs through interpretable substructure analysis but also offers robust predictive performance to support drug development and combination therapy.